This is the second blog post in a series on our Gene Wiki renewal. More details below.
The first funding period focused on crowdsourcing knowledge on human genes in Wikipedia. And that effort has largely been successful. The ~10,000 gene articles are collectively viewed 68 million times per year, and our bot ensures that the “infobox” content is up-to-date and accurate.
But the use of Wikipedia as a resource for biomedical data extends beyond genes. For example, there are currently 5,400 articles on drugs and 5,623 articles on human diseases that are viewed 300 million and 950 million times per year, respectively. Clearly the thirst for this scientific information is high.
This aim expands the scope of the Gene Wiki to include both drugs and diseases. As we did for human genes, we will systematically integrate and aggregate data from existing structured databases, and we will ensure that the data on Wikipedia are coherently presented and current.
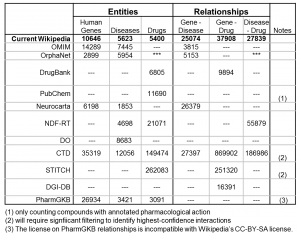 The table at right summarizes a few of the key data resources that will be used for our expanded Gene Wiki. A few details on our plans for each of these biomedical entity types are also presented here:
The table at right summarizes a few of the key data resources that will be used for our expanded Gene Wiki. A few details on our plans for each of these biomedical entity types are also presented here:
- Genes: We will add more complete coverage of SNPs and pathways
- Diseases: We are partnering with the Disease Ontology group to systematically define the universe of human diseases with crosslinks to OMIM, MeSH, UMLS, etc., and also working with the Human Phenotype Ontology group to present the clinical symptoms associated with each disease.
- Drugs: We will primarily use DrugBank to define the universe of human-relevant drugs, augmenting with PubChem and NDF-RT as appropriate.
Perhaps even more exciting than the descriptions of genes, diseases, and drugs individually is better organization of the links between these entities.
- Gene-disease: In collaboration with the Neurocarta team, we will assemble and characterize the links between diseases and the associated genes. These associations can either be directly causative (e.g., “mutation in Gene X causes Disease Y”) or correlative (e.g., “Protein encoded by X is a biomarker for Y”).
- Gene-drug: The most obvious links of this type involve drugs and the proteins that they target, though DGIdb categorizes interactions according to over 40 types (including inhibition, antagonism, agonism, potentiation, and binding).
- Drug-disease: Links between the drugs and their associated diseases will be primarily drawn from NDF-RT, and link types include “may_treat”, “may_prevent”, “may_diagnose”, and “induces”.
Prototypes for all the infoboxes that will be maintained in this effort are being developed online in collaboration with the Wikipedia community at http://en.wikipedia.org/wiki/User:ProteinBoxBot/Phase_3. This aim also heavily involves Wikidata, but I’ll have more to say about this in the discussion of our Aim #3.
This blog post is part of a series of entries on our NIH proposal to continue developing the Gene Wiki. The other posts are here:
Post #0: Introduction
Post #1: Gene Wiki progress report
Post #2: Aim 1: Diseases and drugs (this post)
Post #3: Aim 2: Outreach
Post #4: Aim 3: Centralized Model Organism Database
Post #5: Aim 4: Patient-aligned crowdsourcing
