In BioGPS, there are a number of Intermine–based model organism plugins (and to a lesser extent model organism data sets) which allow users to explore gene expression in organisms typically studied in biomedical research. Model organisms such as mice, rats, flies, worms, zebra fish, etc. have well-annotated genomes and a lot of well-established tools for further exploring and contributing to the knowledgebase around those animals. In contrast, valuable agricultural animals do not have this degree of data, tools, and resource development. This may change as the biomedical and agricultural research domains blur thanks to the movement of medication and infectious disease from farm animals into humans. In this spotlight, we’re happy to introduce a new data set that’s been added to BioGPS–the Sheep Gene Expression Atlas. Emily Clark, a researcher and Chancellor’s Fellow from The Roslin Institute, University of Edinburgh, kindly answered our questions.
- In one tweet or less, introduce us to the Sheep Gene Expression Atlas:
A high resolution atlas of gene expression across tissues and cell types in sheep.
- Who is your target audience? How big is the community studying sheep genetics?
Our target audience is the livestock research community, particularly those working on small ruminants. There is a large research community studying sheep genetics with research groups across the globe and an International Sheep Genomics Consortium (ISGC). The project is also a valuable resource for the Functional Annotation of Animal Genomes Consortium (FAANG) and represents the largest RNA-Seq FAANG dataset to date. Sheep are also an important non-human model and we hope the data will be useful for the mammalian genomics community more generally.
- It looks like the academic article on the Sheep Gene Expression Atlas was published a little more than a month ago in PLOS Genetics. How long has the team been working on the atlas before reaching this point?
The sheep gene expression atlas was initiated in 2013, so we have been working on it for approximately 4 years. The first year involved tissue collection then the following years, library preparation and data analysis.
- In your paper, you illustrate the value of the Sheep Gene Expression Atlas by looking at Innate Immunity genes and the advantages of crossbreeding. What other types of research could this atlas contribute to? Antibody development for immunological assays? Prion disease research? Antibiotic use in animal husbandry?
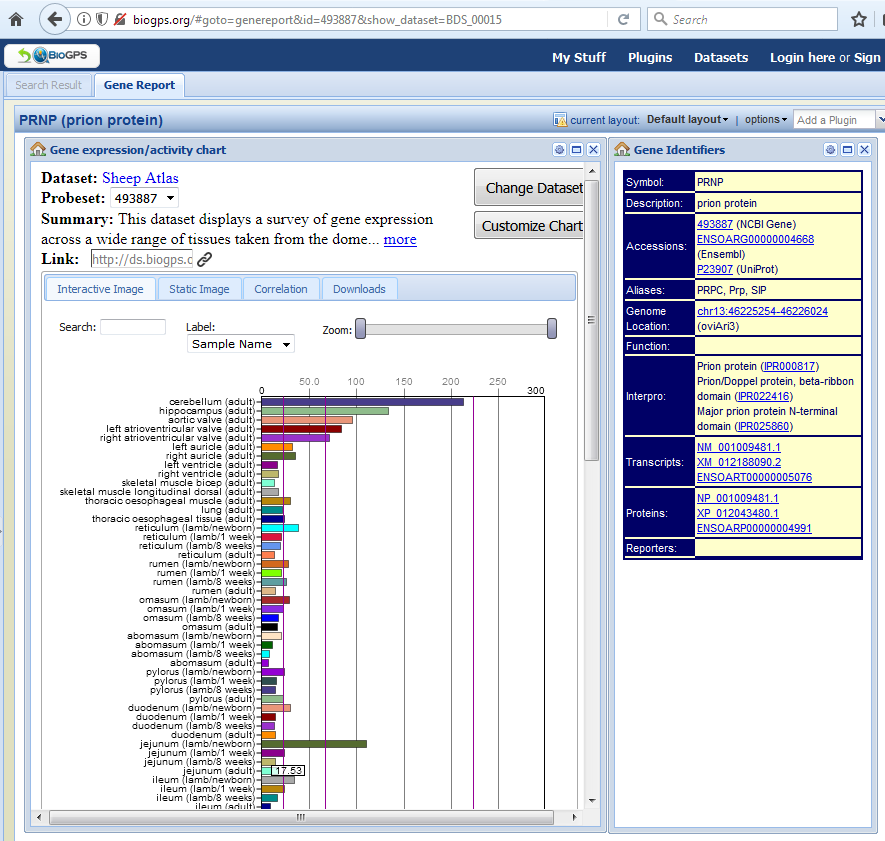
We hope that the atlas will now be used by researchers working in livestock genetics and genomics to link genotype to phenotype. It has potential uses in identifying targets for novel therapeutics, some of the dataset from the sheep expression atlas project has been used to identify genes relevant to resistance to mastitis, for example (Banos et al. 2017 The Genomic Architecture of Mastitis Resistance, BMC Genomics). Researchers at the Roslin Institute, interested in prion disease, are also looking at the expression of the gene PRNP (prion protein) across tissues using the sheep atlas dataset. The scale and scope of the dataset is such that it should contribute and provide information for multiple research projects and different fields in sheep but also other ruminants and livestock.
- Who is the team behind the Sheep Gene Expression Atlas?
The sheep atlas project was led by David Hume and Alan Archibald who initiated the work. It was coordinated by Emily Clark, with bioinformatic support from Stephen Bush. The project involved a large team of people for sample collection at The Roslin Institute including farm technicians who also managed the animals for the project. We are also very grateful to Chunlei Wu and Cyrus Afrasiabi for their help making the sheep atlas dataset visualisable on the BioGPS platform.
- What is in store for the Sheep Gene Expression Atlas?
Next we hope to use the data set for a global analysis of allele specific expression across tissues and cell types in sheep and we also have a comparative analysis of gene expression from a smaller subset of tissues in goat which we hope to release soon.
Thanks to Emily Clark, and the rest of the the Sheep Gene Expression Atlas team, for sharing their high resolution Sheep Gene Expression Atlas with BioGPS. If you use the Sheep Gene Expression Atlas data set in your research, be sure to cite their publication:
To search for your favorite genes in the Sheep Gene Expression Atlas, visit the sheep-specific portal at: http://biogps.org/sheepatlas/#goto=welcome

Trackbacks/Pingbacks