News
We have been accepted as a mentoring organization for GSOC13 !!!
See our gsoc project page for the application template, scroll down for the ideas list, and don’t forget to get on our discussion forum.
Welcome to the “Crowdsourcing Biology” Google Summer of Code summary page!
“Google Summer of Code (GSoC) is a global program that offers student developers stipends to write code for various open source projects. Google will be working with several open source, free software, and technology-related groups to identify and fund several projects over a three month period. Since its inception in 2005, the program has brought together thousands of students and mentors from over 100 countries worldwide, all for the love of code.” — Google GSoC
Although we are a relatively new organization to the GSoC program, our lab has a long history of participating and promoting collaborative science through “crowdsourcing” and open source code development. More info is below on our Gene Wiki initiative, our BioGPS gene portal, and our emerging emphasis on games for biological research! Curious? Get in touch with us on the discussion forum!
Meet the mentors
- Andrew Su, Ph.D. — Associate Professor, overall lab head
- Chunlei Wu, Ph.D. — Associate Professor, BioGPS team leader
- Benjamin Good, Ph.D. — Senior Staff Scientist, Gene Wiki team leader and games team leader
- Salvatore Loguercio, Ph.D. — Postdoctoral associate, Gene Wiki team and games team
- Katie Fisch, Ph.D. — Postdoctoral associate, bioinformatics, statistics
- Max Nanis — Research Programmer, Gene Wiki, Games and BioGPS
- Erik Clarke — Research Programmer, Gene Wiki team
- Ian Macleod — Research Programmer, BioGPS
Feel free to contact any of the mentors above, or join the discussion on the Crowdsourcing Biology Google Group.
Primary Development groups
Gene Wiki
The broad goal of the Gene Wiki initiative is to organize knowledge of human gene function such that scientists and lay people can quickly and easily access it in a gene-centric manner. To accomplish this goal, the Gene Wiki facilitates the development of Wikipedia articles that describe specific human genes. To date, this collection contains more than 10,500 such articles that together contain more than 1.5 million words. These articles are typically the first results returned by Google searches for gene symbols and are collectively accessed more than 50 million times per year. For more information see: http://en.wikipedia.org/wiki/Portal:Gene_Wiki
- Code repository:
- Discussion forum: Crowdsourcing Biology Google Group and http://en.wikipedia.org/wiki/Template_talk:GNF_Protein_box
BioGPS
BioGPS (http://biogps.org) is a free extensible and customizable gene annotation portal, a complete resource for learning about gene and protein function. Users can register any web-based biological resource as a “plugin” in BioGPS, and other users can then use this resource and mix-match with any other desired resources to make their own customized gene annotation page (called a gene-report “layout” in BioGPS). BioGPS receives millions of hits per year, and is freely and publicly available at biogps.org.
- Code repository: https://bitbucket.org/sulab/biogps_core (NOTE: historical code repository not available due to some licensed non-open-source components. Replacing these components with better and open-source versions is one goal of our GSoC application.)
- Discussion forum: Crowdsourcing Biology Google Group and http://groups.google.com/group/biogps
Biological games
While the Gene Wiki and BioGPS have successfully tapped into the wisdom of the crowds for scientific applications, we believe that these efforts are only leveraging a small fraction of the human potential that crowdsourcing offers. In the next phase of our research, we plan to expand the scope and breadth of crowdsourcing science by creating computer games for biology. In this line of work we will build on the successes of Luis Von Ahn and colleagues at CMU (see http://www.gwap.com) as well as the Foldit team from the University of Washington (see http://fold.it) to devise and implement a whole new series of games that will advance biomedical research. Our most recent effort in this area is The Cure, a game that helps generate better predictors of breast cancer prognosis. The discussion forum for each of the games is the Crowdsourcing Biology Google Group and the games will be distributed via http://genegames.org.
Games developed or in development include:
- The Cure, a card game to capture gene sets associated with breast cancer prognosis. (Also integrates machine learning)
- Code repository: https://bitbucket.org/sulab/thecure
- Dizeez, a quiz game to comprehensively catalog links between genes and diseases.
- Code repository: https://bitbucket.org/sulab/dizeez
- GenESP, a game of biological Extra Sensory Perception. For generating gene annotations.
- Code repository: https://bitbucket.org/sulab/genesp
Current Ideas List (2013)
Ideas for GSoC 2013 projects in Crowdsourcing Biology
- Idea 0. Create your own!
- Idea 1. Gene Wiki. Convert Gene Wiki bot to write to WikiData
- Idea 2. Games. Develop a new version of the game Dizeez as a Facebook application
- Idea 3. BioGPS. New gene query interface (backed by MyGene.info v2 API)
- Idea 4. BioGPS. Advanced gene annotation tools: ID mapping and user table annotation.
- Idea 5. BioGPS. Gene report layout “sharing by link” button
- Idea 6. Games. Add interactive decision tree builder for the The Cure
- Idea 7. BioGPS. Add plugin to display content from Semantic BioGPS.
Idea 0. Create your own!
Feel free to propose your own idea. As long as it fits within the general space of ‘crowdsourcing biology’ we would be thrilled to consider it. Please be detailed about what the specific goal would be, why it is important and how you plan to achieve it.
Idea 1. Convert Gene Wiki bot to write to WikiData
Background. Code developed by our group helps to maintain more than 10,000 articles about human genes on Wikipedia. All of the page elements highlighted in orange below are generated by a program that gathers information from trusted public databases (accessed via MyGene.info) and inserts it into the right structure at Wikipedia. Collectively the articles are viewed more than 50 million times a year.
Wikidata is a brand new development at Wikipedia. Its goal is to create a free database that anyone can edit. The aim is to collect all of the “structured data”, the elements of infoboxes for example, into a central, queriable database. Articles on Wikipedia can query this database to access and then render the content much like other web applications. In addition, external web applications can dynamically query this content to enhance their own content. The goal of this project will be to migrate the data that we currently store in template pages in Wikipedia into the wikidata database.
For example, we now get data from mygene.info and add it to templates like this one for the gene Reelin . We would like to move that data instead into the wikidata page associated with Reelin. This will allow us to query the database, others to edit the database, etc. in a much cleaner fashion than we can accomplish working inside of the Wikipedia article/template text directly. This will also form the basis for adding new structured content about genes into Wikipedia such as explicit connections between genes and disease.
Specifics. Convert the pygenewikibot to deposit the data used to populate the gene articles into WikiData. (see issue tracker). Add visualization elements to one or both of the gene wiki (Wikipedia) and GeneWiki+ that display the information from wikidata.
You should consider this project if: you know Python, you are excited about working to make Wikipedia better, and you want your work to be used by millions of people every day.
Possible mentors: Ben, Sal, Max, Andrew
References: Molecular Biology task force for wikidata, a repo for wikidata bot code, gene wiki portal
Idea 2. Develop a new version of the game Dizeez as a Facebook application
Background. Dizeez is a simple quiz game about connections between genes and diseases. Players learn about gene-disease connections through game play. In addition, the system learns new connections suggested by many different players – thus providing a mechanisms to crowdsource the creation of a new gene-disease database. Building on some early success with a simple Dizeez webapp, this project will bring this game into the Facebook context – hopefully garnering many more players, creating a more enjoyable player experience, and producing better data.
Suggested implementation: Facebook Canvas application
- Built as a single page javascript application
- Marionette (w/ Backbone) using RequireJS
- API to sync game play results and get back new gene & disease options.
- FB Integration
- Score API to challenge players in your social graph
- Achievements API to promote level and game advancements
- Notifications to get others to join you
- Dynamic disease choice answer compositions
Mentors: Max
Idea 3. BioGPS. New gene query interface (backed by MyGene.info v2 API)
MyGene.info v2 gene query API expands both data and query capabilities over the current APIs. We designed a new and clean query interface to utilize these new APIs. Here are a few highlights (with mock-ups provided when necessary):
- By default, it supports a single line query input with a rich query syntax (as provided by MyGene.info API) (mockup)
- A gene query result page to show the list of hits returned from query API. (mockup)
- Support Advanced query mode for multi-line queries (e.g. using a “area” element instead of “input”)
- mode 1: auto-switch to multi-line query mode when users paste into a multi-line text into default input box.
- mode 2: switch to multi-line query mode by a link or button. if mode 1 is hard to implement, this will be the fallback option.
- Besides default queries on genes, BioGPS also support queries on other entities, like plugins, datasets, genelists. It will be nice to implement a “Google-Drive” style query options/filters (see “images” filter below)
Refer to this post for setting up a BioGPS dev environment. Also note that, if preferred, this project can be developed as a standalone application as well (without BioGPS dev environment). The integration with BioGPS can be done in later stage.
Mentors: Chunlei, Andrew
Additional information: http://genomebiology.com/2009/10/11/R130 http://nar.oxfordjournals.org/content/41/D1/D561.short
Idea 4. BioGPS. Advanced gene annotation tools: ID mapping and User table annotation.
Idea 3 above implements the general query interface for BioGPS. Besides that, we also want to implement two commonly used gene annotation features:
- ID mapping Users can copy/paste or load (from a file) a list of gene identifiers, and then specify one or multiple identifier types (Entrez GeneID, RefSeq ID, Uniprot ID, etc), the fields they want to output (the default will be “symbol”, “name”, can be any fields supported by MyGene.info API). The output will be a table of matching genes with user-specified output fields.
- User table annotation This is tightly related with “ID mapping”. The difference is that users will load a table from a file (tsv, csv, xlsx, etc), then specify which column is the “identifer” column they want to query for the gene annotations. They will then specify the identifier types and output fields as well. The output fields will be inserted into users’ table as additional columns. A “download” link will be provided for users to download the entire annotated table.
Mentors: Chunlei, Andrew
Additional information: http://genomebiology.com/2009/10/11/R130 http://nar.oxfordjournals.org/content/41/D1/D561.short
Idea 5.������������������������������BioGPS. Gene report layout “sharing by link” button
BioGPS’s gene report page (like this one) is the core landing page for our users, which displays variety of gene-centric resources users picked from our plugin library. We use the concept of “layout” (or “gene report layout”) to represent the view of users’ customized gene report page, which essentially contains the information of plugins identities, window poistions/sizes, etc. This is an example layout. We would like to implement a layout-sharing feature described as the following:
- It works similar to google map’s sharing button below, we will put a “sharing by link” button in gene report page.
- When the button was clicked, a “long” url is generated, which contains all information to re-produce user’s current gene report view.
- Allow users to share the link via common social network (twitter, facebook, etc.)
- Allow to generate a persistent “short-url��� version of the long one (http://biogps.org/view/qMjXHuaG). Another idea is to delegate this feature to external “url shortener” service.
Refer to this post for setting up a BioGPS dev environment.
Mentors: Chunlei, Andrew
Additional information: http://genomebiology.com/2009/10/11/R130 http://nar.oxfordjournals.org/content/41/D1/D561.short
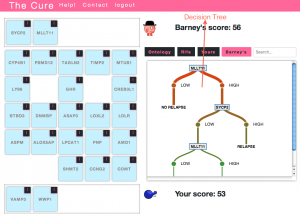
Idea 6. Games. Add interactive decision tree builder to The Cure
Background. The Cure is a game related to machine learning and classification in biological domains (especially breast cancer). In the current implementation, a decision tree is automatically constructed, tested and rendered by the system based only on the player picking which variables (genes) to use to build it. Idea. Create a new game interface that lets the players interactively construct the decision trees based on any number of genes of their choosing. This would basically be an ‘expert mode’ that gave players full control. The trees would be evaluated (server-side) based on how well they classified the training set, how big they were (smaller is better), and how biologically meaningful the group of genes in the tree is.
See repo.
Suggested implementation. D3 javascript.
Mentors: Max, Ben
Idea 7. BioGPS. Add plugin to display content from Semantic BioGPS.
Background: The broad aim of this project is to enable information extraction from all of the websites (plugins) accessible via biogps.org . Specifically, we are attempting to gather gene annotations. ����For example triples like: �������������Gene X plays a role in Alzheimer’s Disease”. To extract this knowledge, we have a Web interface that allows users to visually construct HTML parsers specific to each BioGPS plugin (that end up being Xpath statements). There is also code that runs these parsers over large volumes of gene-centric pages and stores the extracted content in a MongoDB database. (See https://bitbucket.org/sulab/semantic-biogps)
Idea: For this year’s GSoC, we propose the development of a Semantic BioGPS display plugin. This would query the data extracted from all of the annotated plugins and display it in the context of BioGPS. The data to be displayed would be in a triple structure with the gene as the subject and could have a large variety of properties (predicates) and object types. While this interface is the end goal, the project would also need to improve on the current annotation authoring pipeline to ensure that it is robust enough to support a public interface. This will likely involve refinements to the annotation authoring interface and regularly scheduled crawls of the biogps plugin library. Because of the nature of the task (parsing structured data from semi-structured and transient HTML) robust error-handling is vital to consider at every step.
Possible Mentors: Chunlei, Andrew, Ben, Max
Possible interface dev tools: roll your own html/javascript, Exhibit, D3, Cytoscapeweb
A demo site for semantic BioGPS is live here: http://54.244.135.254/
Additional information: http://genomebiology.com/2009/10/11/R130 http://nar.oxfordjournals.org/content/41/D1/D561.short