Finally, biomedical researchers have a single tool that can compare locations and epidemic curves, access datasets, find doubling rates, and search for publications. Outbreak.info standardizes and aggregates COVID-19 and SARS-CoV-2 data for easy access1.
Outbreak.info is a searchable, open repository of global COVID-19 and SARS-CoV-2 data that is updated daily. It is a central hub of all SARS-CoV-2 and COVID-19 information with open access to resources, including publications, preprints, datasets, protocols, and clinical trials. The interactive data dashboards allow for quick comparison of countries, states, counties, and metro areas. And like any good open source tool, all of this data can be downloaded and/or accessed through our API.
Let’s examine four ways researchers can use outbreak.info’s open source data visualizations to get a snapshot of the world today and quickly draw insights.
#1 Use the searchable interface to find new resources in a matter of seconds.
Efficient open data sharing is important. With so much international collaboration and an inundation of open science, how do researchers manage what the WHO has coined the “infodemic”2? Outbreak.info recognizes that researchers need a searchable database and offers that solution. Outbreak.info promotes international sharing of COVID-19 and SARS-CoV-2 literature with an aggregated, searchable, free online collection of scholarly articles, preprints, peer reviewed journals. The repository is updated daily – start your search or browse the newest additions.
For example, in a medRxiv preprint published this week, researchers apply physical and mathematical principles with game theory to highlight the difference in timelines that countries may experience if they follow non-cooperate models vs. strong cooperative models3: https://outbreak.info/resources/2020.12.01.20242099
#2 Track the global coronavirus pandemic with a live dashboard.
The interactive global map allows researchers to view countries at a glance, compare US numbers, and spot critical trends.
For example, we can use the tool to find locations where case numbers have been lower but are increasing. We can select “two week change in cases per day per capita” to find countries that have experienced the greatest increase in cases over two weeks per 100,000 people.
Here we see that Belize is currently experiencing the greatest increase with 105 more cases per 100,000 people (over a four day average) compared to two weeks ago.
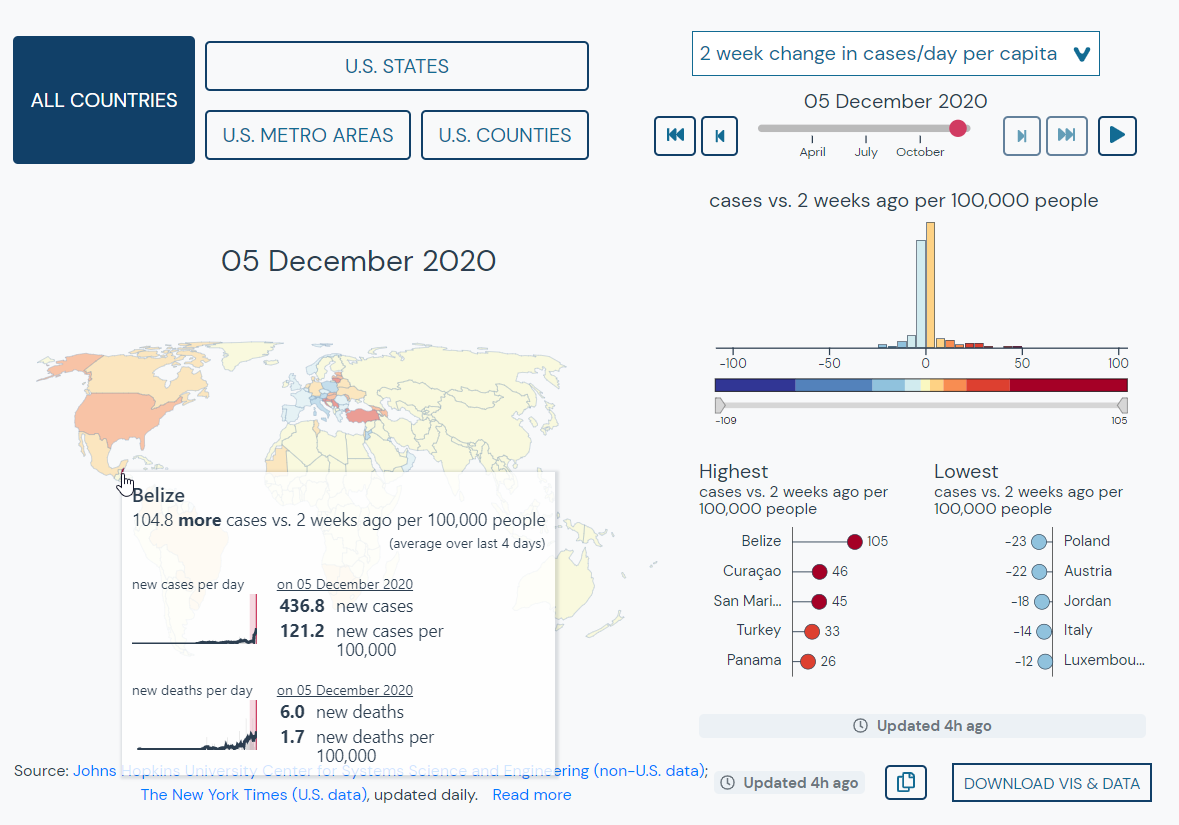
#3 Find locations that are similar in population, cases, or deaths.
We can use Compare Locations to find places that are similar in population, new cases, deaths, etc. To continue the example, we can find countries, states, and territories that are similar to Belize today in new cases per capita with a simple search.
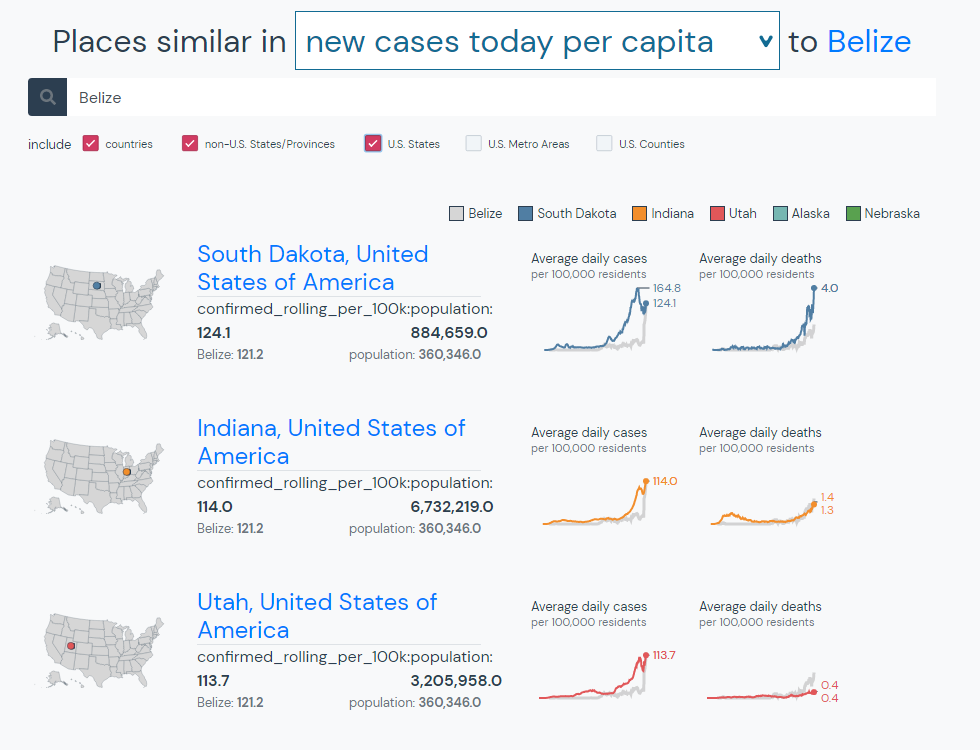
#4 Compare epidemic curves for locations around the world over time.
This open source tool can be used to compare different locations around the world by simply overlaying epidemic curves for different locations and normalizing to population.
We can use this tool to see how Belize has been doing over time.
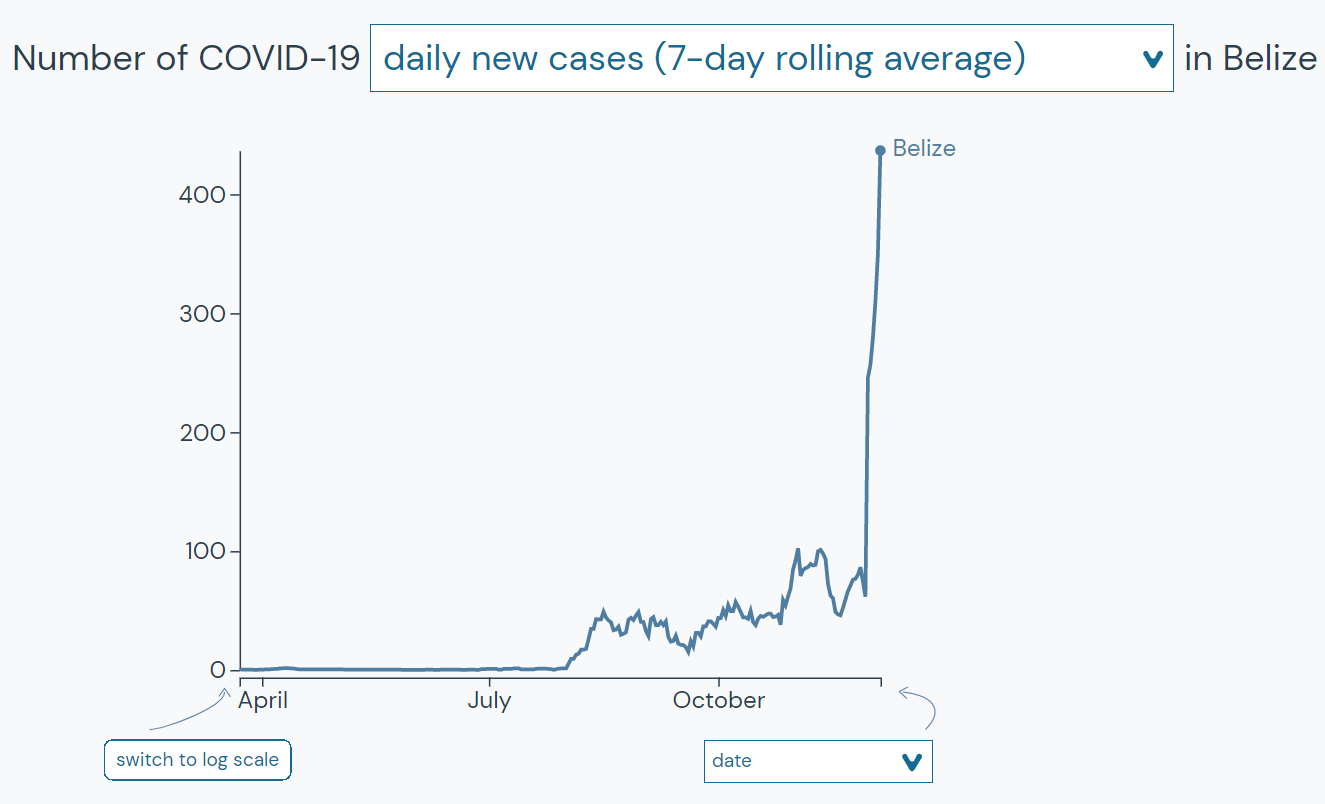
We can also use this tool to see how locations that we identified as being similar to Belize today have been doing over time.
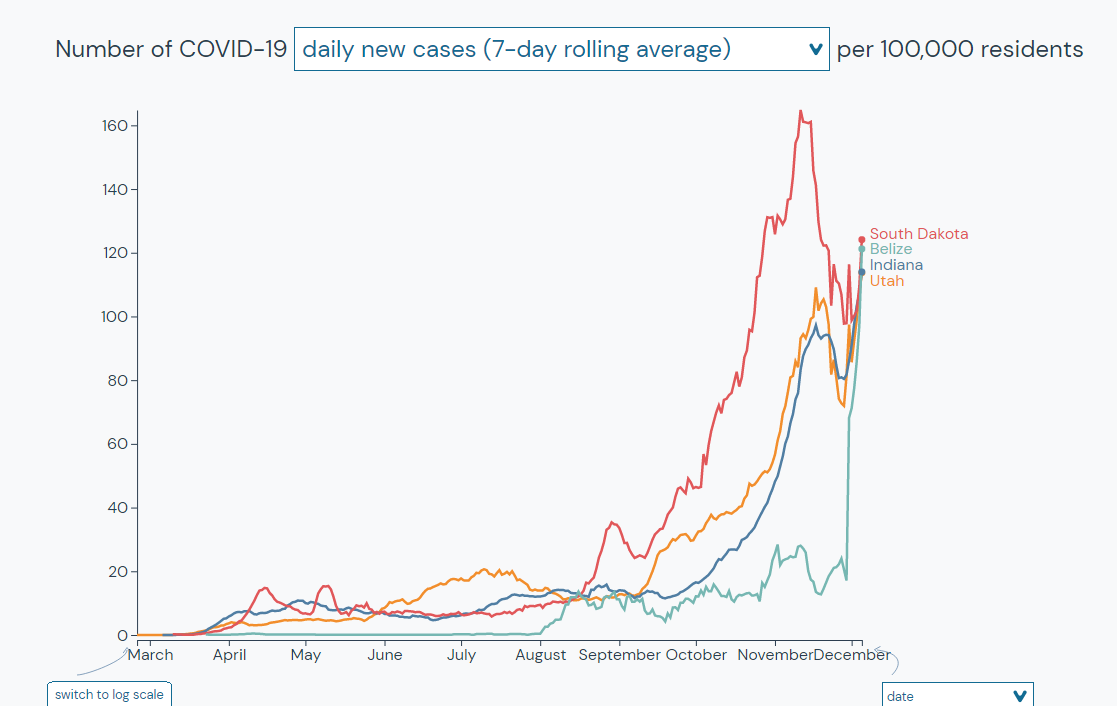
Outbreak.info is the centralized hub of COVID-19 and SARS-CoV-2 resources that epidemiologists and biomedical researchers have been asking for. This publicly available coronavirus resource center provides needed accessibility for researchers around the globe.
Read more about outbreak.info resources.
Source
[1] Hughes, Laura D.; Gangavarapu, Karthik; Alkuzweny, Manar; Cano, Marco; Mullen, Julia; Rush, Benjamin; Tsueng, Ginger; Zhou, Jerry; Andersen, Kristian G.; Wu, Chunlei; Su, Andrew I. outbreak.info. Available online: https://outbreak.info/ (2020)
[2] Zarocostas, John. How to fight an infodemic. The Lancet. 2020.02.27.0140-6736;
doi: https://doi.org/10.1016/S0140-6736(20)30461-X
[3] Garcia de Alcaniz, Javier G.; Lopez-Rodas Sr., Victoria; Costas Sr., Eduardo; et al. Is the end near? When the different countries will surmount COVID-19 pandemic: new approach applying physical, mathematical and game theory models. medRxiv. 2020.12.01.20242099; doi: https://doi.org/10.1101/2020.12.01.20242099
