Crowdsourcing is clearly a theme in our lab, but one flavor of crowdsourcing that we hadn’t experimented with extensively is the use of prize-based contests. Sites like TopCoder, Innocentive, and Kaggle have shown that contests can be a successful mechanism for crowdsourcing in diverse communities of expertise.
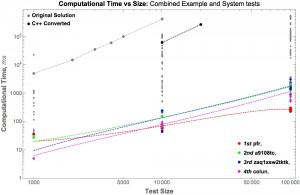
One excellent demonstration of contests for biomedical applications came from Karim Lakhani and his colleagues (published in Nature Biotechnology in 2013). They used a TopCoder contest to optimize a bioinformatics algorithm for assigning V, D, and J gene usage from mature antibody sequences.
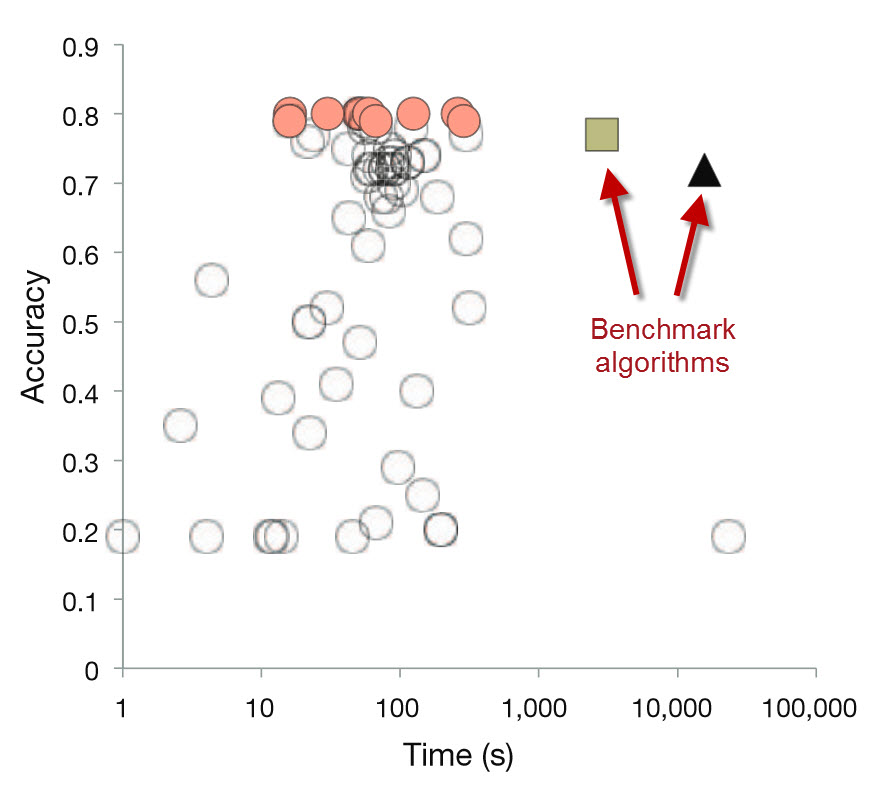
The TopCoder contest resulted in many solutions that were more accurate and 100- to 1000-fold faster.
Relative to the benchmark algorithms, the TopCoder contestants were able to improve the efficiency by 2-3 orders of magnitude while simultaneously increasing the accuracy of the results.
Contest duration: Two weeks
Prize Pool: $6000
Since I knew Karim from other interactions (including a visit to the Pentagon), we reached out to him to set up two new contests for tough challenges we were facing.
Contest #1: Antibody clustering
In collaboration with the Scripps Center for HIV/AIDS Vaccine Immunology and Immunogen Discovery, we were interested in optimizing another antibody sequence analysis problem. Specifically, given the sequences for an entire antibody repertoire, we wanted to perform clonal lineage assignments. The basic goal was to group these antibody sequences according to the naive B cell from which they originated.
Brian Briney had created Clonify, an initial version of this algorithm based on sequence-based clustering. But while sequencing an antibody repertoire might result in millions of sequences, CPU and memory constraints meant that we were nowhere near handling that size of experiment. After a little bit of Python optimization from Chunlei, Clonify could classify up to 100k sequences on a high-memory server (150 GB) in 1.7 hours. Using this as the baseline, we posed this optimization contest to the TopCoder community.
The top scoring solution successfully clusters 2.3M sequences in ~30 seconds and requires only 1.1 Gb of RAM. That solution represents a > 20-fold improvement in capacity, 10,000-fold improvement in speed, and > 10-fold improvement in memory efficiency.
Contest duration: 10 days
Prize Pool: $7500
Contest #2: Named Entity Recognition in biomedical text
As part of our initiative to create a Network of BioThings, we have been developing methods to use crowdsourcing to perform Named Entity Resolution (NER) in text. Previously, we used Amazon Mechanical Turk (AMT) to reproduce the NCBI Disease Corpus, a collection of 793 PubMed abstracts containing 6892 mentions of “disease concepts”. In short, we distributed this task to a crowd of 145 workers, where each abstract was viewed by 15 individuals. We found that in aggregate, the answers from those 15 workers was on par with the professional biocurators who originally created the corpus.
Although the crowd data proved to be quite reliable on their own, we next wanted to test whether they could be used to help train a better computational NER tool. For that, we worked in collaboration with Zhiyong Lu and Robert Leaman at NCBI. Zhiyong’s group was responsible for creating the NCBI Disease Corpus, and Robert created the BANNER system for NER during his graduate work.
Normally, BANNER is trained based on a set of gold standards. Those gold standards are strictly binary — a given word or phrase is annotated as a disease mention, or it isn’t. We hypothesized that the quantitative agreement levels in the AMT data (as well as the longitudinal data for each worker over many abstracts) could be additional information that could be used to train BANNER. Therefore, we set up a second contest to improve BANNER based on AMT data.
While we’re still analyzing these results, our preliminary look suggests another successful contest!
Contest duration: 8 days
Prize Pool: $30860 (several contest formats)
Next steps
Our two experiments above represent very different types of contests, and in both cases it appears that we achieved a very successful result. As we contemplated the next contests we might think about running, I offered this tweet:
suggestions on bioinformatics algorithms where 10-100x speed up would be game changing?
— Andrew Su (@andrewsu) March 16, 2015
Yes, perhaps a tad hyperbolic, but I thought it would get the discussion started (and I did get a few replies via Twitter). Any other ideas to share?