Dizeez @ Future of Genetic Medicine V
Recently, we have added several new features to the Dizeez game (previously described here). In particular, the game now allows players to select a specific area of biology (for example, by disease or protein family) that best matches their expertise. Also, a “recap” functionality has been added at the end of the game, that shows supporting evidence (in Gene Wiki and PubMed GeneRIFs) for each gene-disease association played. Users can review the game log and even suggest new evidences.
To ‘live test’ the new features, we brought the game to The Future of Genomic Medicine V conference held March 1 to 2 2012 at the Scripps Institution of Oceanography in La Jolla. In two days, we had 189 games played to completion by over 60 unique individuals – 22 registered users played 143 out of 189 games. Overall, players provided 2026 guesses across 1791 unique gene-disease assertions.
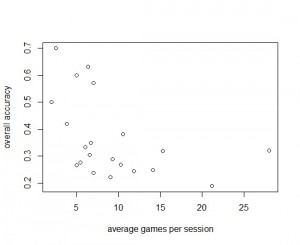
Notably, almost half of our registered users provided ten or more guesses with overall accuracy higher than 30%. Perhaps not surprisingly, overall accuracy seems to correlate with the amount of time spent on each association (Ben would agree here) – so throwing random guesses at it might be not a good strategy for players. Equally importantly, this observation reveals a useful filtering metric for our downstream data mining.
As before, we mined the game logs for novel gene annotations, i.e. gene-disease links well established in the literature, but not yet mirrored as structured annotations. Among the gene-disease assertions provided most often by game players, we had 13 associations occurring 4 or more times. Since all of these associations were previously known via Gene Wiki and/or OMIM, they just provided a positive control.
After removing all gene-disease links already annotated in Gene Wiki/OMIM/PharmGKB, there were 18 assertions that were provided by players two or more times. We quickly ranked them using the Normalized Medline Distance (NMD) as a proxy for gene and disease co-occurrence in Pubmed articles. Results could easily be verified by a quick literature search. The top five associations are summarized here:
| Gene Symbol | Disease name | Disease Ontology ID | NMD | PubMed support |
|---|---|---|---|---|
| ABCB5 | acute myeloid leukemia | DOID:9119 | 0.75 | 22044138, 19477512, 19394083 |
| HOXB7 | leukemia | DOID:1059 | 0.83 | 21183939, 20672360 |
| SULF1 | carcinoma | DOID:305 | 0.83 | 21104785, 21851062 |
| ALPP | retinoblastoma | DOID:768 | 0.87 | 15172750 |
| FOXM1 | Melanoma | DOID:1909 | 0.89 | 22280162, 22094256 |
Interestingly, almost all the publications listed above are very recent (2010-12).
Once again, the game proved useful in identifying novel gene-disease annotations in a short period of time based on the voluntary contributions of a self-selected group of players.

