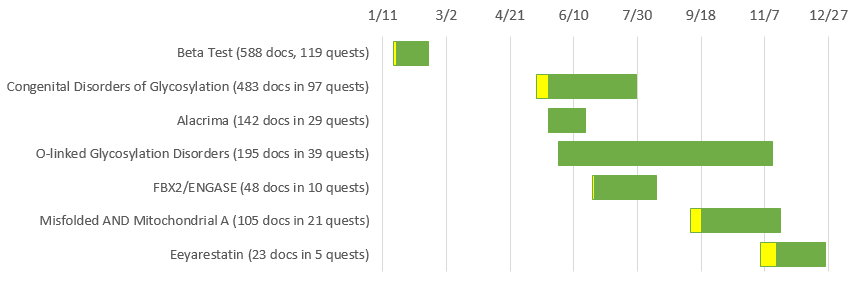
It’s the last day of the year and the perfect time to review the huge strides that Mark2Cure made in 2015. Around this time last year, we conducted an extremely limited beta test just to check and see if this could be done and after 3-5 people completed a set of about 30 documents, we made some minor adjustments and launched a public beta test of Mark2Cure. The beta test began on January 19th and (thanks to researchers and rare disease advocates on twitter, the NGLY1 community, an article in the Union Tribune by Bradley Fikes, and over 230 citizen scientists) was completed on February 16th.
The beta test introduced us to some wonderful participants who provided feedback and guidance as we developed Mark2Cure for the NGLY1 campaign, and an experienced news manager who provided guidance on press outreach. On May 22nd, as Andrew and Matt Might both delivered exciting presentations at the 2015 Stanford Big Data in Biomedicine conference, Mark2Cure was launched along with an excellent article in Science Insider thanks to the amazing Esther Landhuis.
Since the launch of Mark2Cure for the NGLY1 campaign, we’ve had over 469 users join us (though only 392 of them finished all the tutorials.) Thanks to those of you who joined us and contributed, a total of six document sets have been completed so far, and the two currently available doc sets are over 50% complete. Over 437,000 annotations have been submitted for the 1,558 documents that have been made available so far. Together with the submission from the Beta study, there were over HALF A MILLION annotations submitted from over 500 users this year.
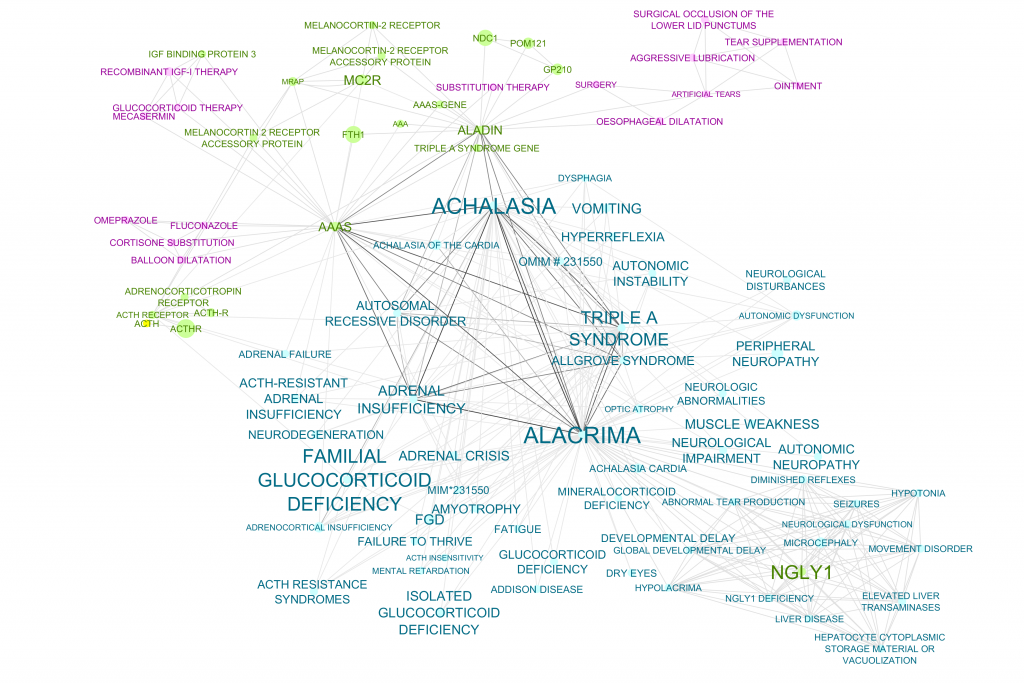
The completion of the Alacrima doc set allowed us to build a network, revealing ACTH as a potential treatment, and we were excited to hear that the Might’s had actually tried this treatment before.
On twitter, we’ve grown from having 72 followers at the beginning of the year to 750 followers. According to twitter analytics, our tweets had 976 retweets, 155 replies, 953 likes, 545 profile clicks, and 1253 url clicks. Some of the tweeps we engaged with most were
@geordiesteph3, @primary_immune1, @mattrussell_PhD, @matteovinci, @HashisNme, @mattmight, @bertrandmight, @rvidal, @evoMRI, @BethNanis, @ObiGriffith, @McDawg, @NewScience101, @jamesian, @globalgenes, @AKUSociety, @SavingCase, @CurityMD, @ERamosSD, @katatonikkat, @bobigelow, @BabyBoomerWritr, @coopsciscoop.
Last and most importantly, we’d like to thank all of the Mark2Curators who have joined us this year. Your criticism, suggestions, complaints, praise, questions, and other feedback helped improve Mark2Cure. We can’t wait to hear what you have to say about some of the new features we’re expecting to roll out in 2016.




Trackbacks/Pingbacks