Last week, I read The Professor and the Madman, a fascinating account by Simon Winchester about Dr. William Chester Minor and the creation of the Oxford English Dictionary. Dr. Minor was one of the greatest volunteer contributors to the OED, and he did it all from his cell at the Broadmoor Asylum for the Criminally Insane. Dr. Minor, who served as a Union Army surgeon in the American Civil War, suffered from schizophrenia (potentially exacerbated by PTSD) and committed murder in a psychotic rage, which landed him at Broadmoor for almost 40 years. The book was beautifully written and speckled with various definitions and etymologies, and for a brief moment or two, I totally understood why Ammon Shea would commit to the ridiculous endeavor of reading the entire OED!
Then, over the weekend, after a long day with our families, my two best friends and I decided to unwind and watch the movie Bad Words. I can’t say I’d recommend the movie as fervently as I do the book, but it wasn’t bad, and it sustained my newfound obsession with words and their histories.
And now with that almost completely irrelevant introduction out of the way, I can begin my post, beginning with a word whose definition and history are so much more than expected from its name:
Interleukin (ĭn′tər-lōō′kĭn) noun. Any of a group of secreted proteins, important in the immune response and involved in the development, differentiation, and movement of lymphocytes. <Latin, inter- (between) + Greek, leukocytes from leukos (white) + kytos (cell)>
The term “interleukin” was coined in 1979 at the Second International Lymphokine Workshop in Ermatingen, Switzerland, to rename Interleukin I (formerly known as lymphocyte-activating factor, mitogenic protein, helper peak-1, T cell-replacing factor III, macrophage-derived T cell-replacing factor, B cell-activating favor, and B cell differentiation factor) and Interleukin II (formerly known as thymocyte stimulating factor, thymocyte mitogenic factor, T cell growth factor, co-stimulator, killer cell helper factor, and secondary cytotoxic T cell-inducing factor).1 At present, 36 interleukins have been identified. They are released by several different cell types (not just leukocytes) and have an ever growing-repertoire of functions. So, even after the very necessary consolidation of terminology in 1979, the interleukins continue to outshine their name. So, in a way, we might as well just call them something like C11ORF24 (see Melissa’s post from last week), because their functions cannot be contained in just a few syllables.
Interleukin 6 reminds me of party streamers (from Wikipedia)
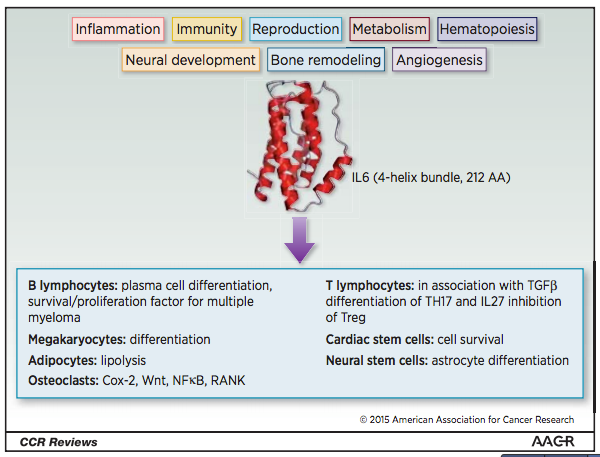
Our new Gene-of-the-Week, IL6, codes for Interleukin 6. IL6 is produced by T cells and macrophages, as well as epithelial, muscle, stromal, and other cells. IL6 binds to a membrane receptor, IL6R, or a soluble receptor, sIL6R. The IL6/IL6R complex then binds to glycoprotein 130 (gp130), setting off a variety of complex signaling cascades.
As can be expected based on many of our previously featured genes, IL6 has been considered a target for anti-cancer therapies. In this case, however, trials have not proven terribly successful. In 2015 review, Rossi et al. speculate that anti-IL6 treatment may fail due to tumor cell plasticity. Although the treatment might work for some tumor cells, there are several other tumor clones that can grow via non-IL6 pathways, so the tumor will continue to grow.2 Despite its lack of success as an anti-cancer target, however, IL6 seems to be a promising target for autoimmune and inflammatory diseases, like rheumatoid arthritis and systemic lupus erythematosus, which have been a highlight of several of my previous blog posts.3 Anti-IL6 therapies have also been successful in treating Castleman’s disease, a cancer-like condition that involves the overgrowth of lymphoid tissue.4
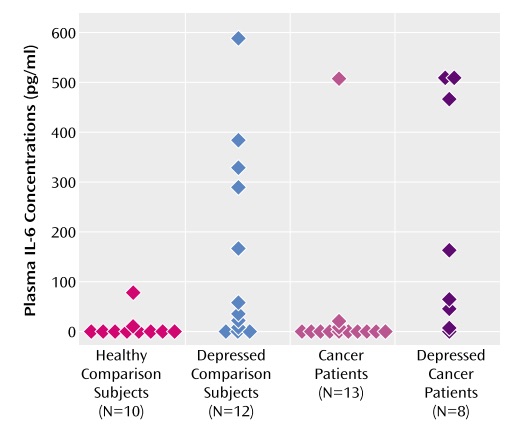
In a small study, Musselman et al. (2001) confirmed that depressed patients have higher plasma IL6 concentrations than healthy controls. Furthermore, they found that depressed patients with cancer have the highest mean plasma IL6 concentrations when compared to patients with depression alone, patients with cancer and no depression, and healthy control subjects. Sample sizes were small, but the study raises interesting questions about the relationship between IL6 levels and rates of depression cancer patients.
For several years, IL-6 high levels have been associated with depression.5 Patients with depression have a significantly higher release of acute-phase proteins, which are regulated by IL6.6 Recently, Yang et al. (2015) reported that serum IL6 levels can be used to predict whether or not a treatment-resistant patient with major depressive disorder will respond to the fast-acting antidepressant effects of ketamine. Patients that responded to ketamine treatment had higher baseline IL6 levels than non-responders and healthy controls. Interestingly, over the course of treatment, IL6 levels decreased. The mechanism by which ketamine acts on depression is currently unknown. It will be interesting to learn how changes in IL6 are related to and possibly involved in ketamine’s mechanism of action.
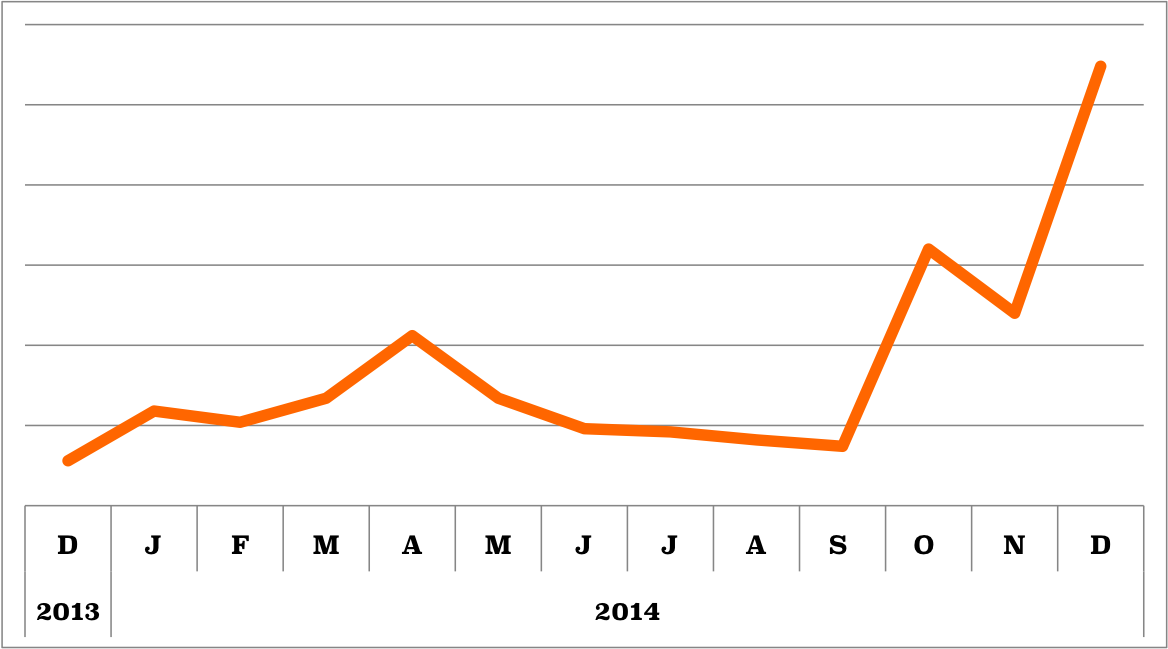
IL-6 is consistently popular among bioGPS users. The recent surge in inquiries may have been prompted by the recent use of anti-IL6 and anti-IL6R drugs to treat inflammatory diseases and major depression.
…Try to fit all of that into a name!
References:
- Aarden, L. A., Brunner, T. K., Cerottini, J.-C., Dayer, J.-M., Ivan, A. L. de W., Dinarello, C. A., Sabato, G. D., et al. (1979). Revised Nomenclature for Antigen-Nonspecific T Cell Proliferation and Helper Factors. Journal of immunology, 123(December), 2928–2929. [↩]
- Rossi, J.-F., Lu, Z.-Y., Jourdan, M., & Klein, B. (2015). Interleukin-6 as a Therapeutic Target. Clinical cancer research : an official journal of the American Association for Cancer Research. doi:10.1158/1078-0432.CCR-14-2291 [↩]
- Singh, J., Beg, S., & Lopez‐Olivo, M. (2010). Tocilizumab for rheumatoid arthritis. The Cochrane Library, (7). [↩]
- Beck, J., Hsu, S., Wijdenes, J., Bataille, R., Klein, B., Vesole, D., Hayden, K., et al. (1994). Brief report: alleviation of systemic manifestations of Castleman’s disease by monoclonal anti-interleukin-6 antibody. New England Journal of Medicine, 330(9), 602–5. [↩]
- Musselman, D. L., Miller, A. H., Porter, M. R., Manatunga, A., Ph, D., Gao, F., Penna, S., et al. (2001). Higher Than Normal Plasma Interleukin-6 Concentrations in Cancer Patients With Depression : Preliminary Findings, (August), 1252–1257. [↩]
- Miller, A. H., Maletic, V., & Raison, C. L. (2009). Inflammation and its discontents: the role of cytokines in the pathophysiology of major depression. Biological psychiatry, 65(9), 732–41. doi:10.1016/j.biopsych.2008.11.029 [↩]