First introduced to the United States in the 1960s, sushi gradually became popular enough, that by the 1980s, American medical journals began warning against the dangers of consuming raw fish.
But this did nothing to stop people from craving sushi, and by 1998, the number of sushi bars in the U.S. had quintupled in just twenty years!1 Today, the California Roll is as ubiquitous, beloved, and as thoroughly “American” as General Tso/Tsao/Gao/Tao’s Chicken. And as a fervent sushi-gobbler myself, I’ll admit that as I write this, there’s about two days’ worth of sashimi currently digesting in my body. But believe it or not, there’s actually some “sushi” inside all of us, all the time—and it’s pretty important.

Unlike these delicious morsels, above, sushi domains are protein domains that only look like real sushi. (Source: Luciano Meirelles, Flickr)
Sushi domains (also called short consensus repeats) are conserved protein domains with a known structure consisting of a protein loop within another loop.2 Many proteins often contain multiple, repeating sushi domains, giving rise to the characteristic resemblance to its delicious namesake. For example, SRPX (descriptively named, “Sushi Repeat Containing Protein, X-linked”) has three sushi domains, which are critical for SRPX’s known functions.
Known primarily as a tumor suppressor gene, SRPX is down-regulated in a variety of malignant cancers, including in neuroendocrine3 and prostate carcinomas.4 Indeed, 30% of SRPX knockout mice develop malignant lymphomas, lung adenocarcinomas, and hepatomas within the first year of life.5 When over-expressed, SRPX is sufficient to suppress tumor formation by enhancing apoptosis; while exact mechanisms remain unknown, it is known that SRPX’s sushi domains are required for this tumor-suppressing activity.678 The sushi domains are thought to act as protein binding regions, though endogenous binding partners (and thus downstream signaling pathways) have yet to be found.

SRPX was originally discovered via its connection to retinitis pigmentosa. This is a photograph of the U.S. Supreme Court Building, modified to show what a patient with retinitis pigmentosa might see. (Source: Wikimedia Commons)
Besides its involvement in cancers, SRPX has also been implicated in a wide range of diseases, including retinitis pigmentosa,9 and more recently, Duchenne muscular dystrophy.10 With retinitis pigmentosa, patients experience a progressive loss of vision, as photoreceptors in the retina degenerate; with Duchenne muscular dystrophy, patients experience gradual muscle loss that eventually results in paralysis, and finally death. Despite the dissimilarities, these rare genetic diseases are occasionally diagnosed in the same patient—mostly because the genes affected in these conditions lie next to each other on the same chromosome!11 In fact, SRPX is located in the same region (Xp21.1) on the X chromosome.12 Therefore, the coincidence of all these conditions could just result from mutations that affect multiple genes in close proximity to each other!
Still, some argue that SRPX could have widespread effects on a number of biological processes. Recent bioinformatics-based analysis has identified another evolutionarily conserved domain of SRPX, suggesting a possible role in peroxide signaling.13 Furthermore, we still don’t know all that much about the other protein domains of SRPX and the exact signaling cascades that they may be involved in. The sushi domains, aside from having a surprisingly apt (and cute!) name, are clearly important for tumor suppression—and perhaps other functions too…
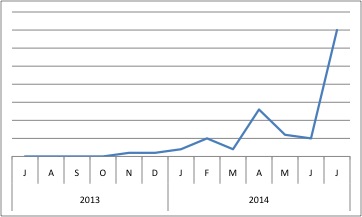
The last known publication involving SRPX was in 2013. Perhaps this increase in popularity (as measured by BioGPS) means that someone’s publishing soon… perhaps we’ll finally know what the sushi domains do!
References:
- “Sushi in America.” Food and Wine. Retrieved 22 August 2014 from http://www.foodandwine.com/articles/sushi-in-america [↩]
- Meindl A, Carvalho MR, Herrmann K, Lorenz B, Achatz H, Lorenz B, Apfelstedt-Sylla E, Wittwer B, Ross M, Meitinger T. (1995) A gene (SRPX) encoding a sushi-repeat-containing protein is deleted in patients with X-linked retinitis pigmentosa. Hum Mol Genet 4(12):2339-46. [↩]
- Pawłowski K, Muszewska A, Lenart A, Szczepińska T, Godzik A, Grynberg M. (2010) A widespread peroxiredoxin-like domain present in tumor suppression- and progression-implicated proteins. BMC Genomics 11:590. [↩]
- Kim CJ, Shimakage M, Kushima R, Mukaisho K, Shinka T, Okada Y, Inoue H. (2003) Down-regulation of drs mRNA in human prostate carcinomas. Hum Pathol 34(7):654-7. [↩]
- Tambe Y, Yoshioka-Yamashita A, Mukaisho K, Haraguchi S, Chano T, Isono T, Kawai T, Suzuki Y, Kushima R, Hattori T, Goto M, Yamada S, Kiso M, Saga Y,Inoue H. (2007) Tumor prone phenotype of mice deficient in a novel apoptosis-inducing gene, drs. Carcinogenesis 28(4):777-84. [↩]
- Pawłowski K, Muszewska A, Lenart A, Szczepińska T, Godzik A, Grynberg M. (2010) A widespread peroxiredoxin-like domain present in tumor suppression- and progression-implicated proteins. BMC Genomics 11:590. [↩]
- Tambe Y, Yoshioka-Yamashita A, Mukaisho K, Haraguchi S, Chano T, Isono T, Kawai T, Suzuki Y, Kushima R, Hattori T, Goto M, Yamada S, Kiso M, Saga Y,Inoue H. (2007) Tumor prone phenotype of mice deficient in a novel apoptosis-inducing gene, drs. Carcinogenesis 28(4):777-84. [↩]
- Kawai T, Suzuki Y, Yamashita A, Inoue H. (2002) Isolation of a novel mouse variant of the drs tumor suppressor gene. Cancer Lett 183(1):79-86. [↩]
- Meindl A, Carvalho MR, Herrmann K, Lorenz B, Achatz H, Lorenz B, Apfelstedt-Sylla E, Wittwer B, Ross M, Meitinger T. (1995) A gene (SRPX) encoding a sushi-repeat-containing protein is deleted in patients with X-linked retinitis pigmentosa. Hum Mol Genet 4(12):2339-46. [↩]
- An HB, Zheng HC, Zhang L, Ma L, Liu ZY. (2013) Partial least squares based identification of Duchenne muscular dystrophy specific genes. J Zhejiang Univ Sci B 14(11):973-82. [↩]
- Watkins CE, Litchfield J, Song E, Jaishankar GB, Misra N, Holla N, Duffourc M, Krishnaswamy G. (2011) Chronic granulomatous disease, the McLeod phenotype and the contiguous gene deletion syndrome–a review. Clin Mol Allergy 9:13. [↩]
- Meindl A, Carvalho MR, Herrmann K, Lorenz B, Achatz H, Lorenz B, Apfelstedt-Sylla E, Wittwer B, Ross M, Meitinger T. (1995) A gene (SRPX) encoding a sushi-repeat-containing protein is deleted in patients with X-linked retinitis pigmentosa. Hum Mol Genet 4(12):2339-46. [↩]
- Pawłowski K, Muszewska A, Lenart A, Szczepińska T, Godzik A, Grynberg M. (2010) A widespread peroxiredoxin-like domain present in tumor suppression- and progression-implicated proteins. BMC Genomics 11:590. [↩]

Trackbacks/Pingbacks