![]() Well, truth be told, the ability for users to register their own plugins in BioGPS has been live for quite a while. The only thing holding up the official announcement has been the creation of the accompanying screencast, which thankfully is now done.
Well, truth be told, the ability for users to register their own plugins in BioGPS has been live for quite a while. The only thing holding up the official announcement has been the creation of the accompanying screencast, which thankfully is now done.
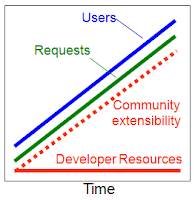 But make no mistake, this is absolutely a critical milestone in our BioGPS development path. Open plugin registration allows any user to add new gene annotation resources into BioGPS. This feature supports one of our two fundamental principles, specifically the principle of community extensibility. By enabling the community of developers to scale with the community of users, we overcome the Law of inevitable stagnation.
But make no mistake, this is absolutely a critical milestone in our BioGPS development path. Open plugin registration allows any user to add new gene annotation resources into BioGPS. This feature supports one of our two fundamental principles, specifically the principle of community extensibility. By enabling the community of developers to scale with the community of users, we overcome the Law of inevitable stagnation.
Of course, this is dependent on developers actually registering their content in BioGPS. But why would people want to contribute to BioGPS? Are we naively relying on the altruism of others? Absolutely not. We’re appealing to the motivating factor for all successful social applications — self-interest.
Consider these advantages of registering your site as a BioGPS plugin:
- You don’t worry about synonym and identifier resolution. You can pick the one “native” identifier you want to use, and BioGPS automatically translates the user query into your identifier of choice.
- Tap into the substantial BioGPS/SymAtlas user base for instant visibility. Combined, we reach a worldwide audience of 50,000+ researchers over 2 million page views per year.
- You’ve probably done the hard work already. Hooking up your site as a BioGPS plugin will likely take five minutes or less, as demonstrated in the screencast above.
- Allow your users to view your data in the context of other gene annotation information. Using just the 93 public plugins that are already in our plugin library, there are already 1028 unique combinations in which to display plugins in BioGPS!
(Oh, and we’ll send some BioGPS swag to the first ten people who register plugins…)
